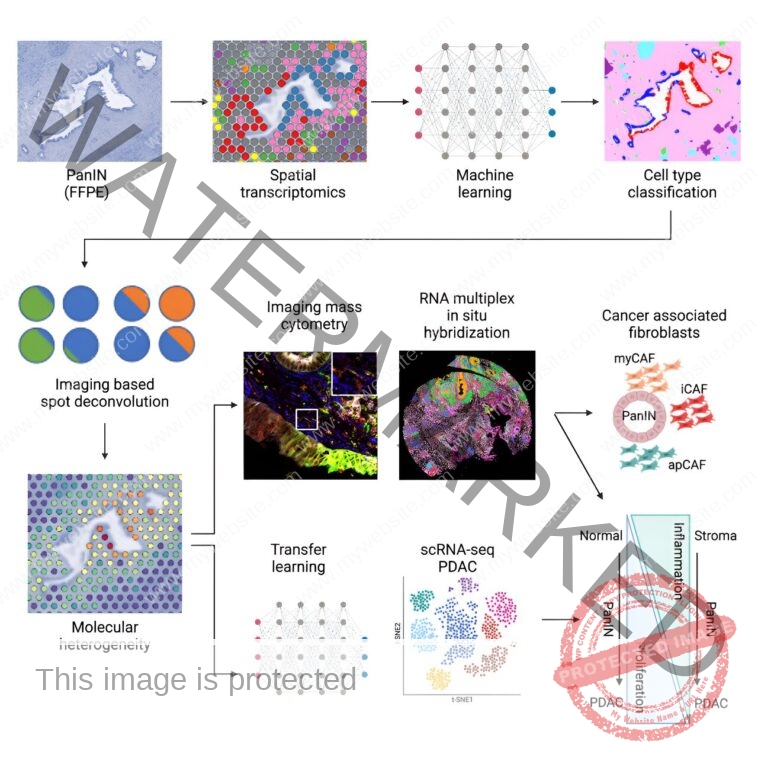
Using a brand new workflow that integrates spatial transcriptomics and machine studying for imaging evaluation and integration with single-cell datasets, researchers on the Johns Hopkins Kimmel Cancer Center have recognized novel molecular and mobile markers within the growth of probably the most aggressive, lethal pancreatic cancers: pancreatic ductal adenocarcinoma (PDAC).
PDAC arises from precancerous lesions within the pancreas. One of those lesion sorts, pancreatic intraepithelial neoplasias (PanINs), can seem within the pancreas years earlier than they progress to invasive most cancers. Because PanINs are so small, they can’t be detected by standard medical imaging assessments.
Previous evaluation strategies, similar to bulk sequencing and single-cell sequencing, can seize gene expression of most cancers cells and different cell sorts within the tumor microenvironment. However, what’s lacking is the spatial relationships amongst a majority of these cells inside and round tumors, says senior research co-author Luciane Kagohara, Ph.D., an assistant professor of oncology on the Johns Hopkins University School of Medicine.
Elana Fertig, Ph.D., professor of oncology, director of the oncology quantitative sciences division, and co-director of the Convergence Institute and the Single-Cell Training and Analysis Center on the Johns Hopkins University School of Medicine, was additionally a senior research co-author.
Using spatial transcriptomics—a method used to measure and map gene expression in a tissue part—the researchers developed a three-way evaluation pipeline to map modifications in gene expression from 9 sufferers in 14 PanINs, together with 5 uncommon, excessive grade PanINs. The machine studying instruments used for imaging analyses (CODA) and for the combination with PDAC single-cell datasets, by the modern multiomics integration strategies CoGAPS (coordinated gene exercise in sample units) and projectR, had been developed in earlier research at Johns Hopkins.
The investigators have made their information and code out there to different researchers to uncover additional insights into pancreatic most cancers and, by open-source instruments, to adapt the evaluation strategies for spatial transcriptomics for biomedical analysis at giant.
Results of their work, printed Aug. 7 in Cell Systems, present new insights into the gene expression and spatial distribution of various kinds of cells within the precancerous setting round PanINs. The findings are key to understanding how PanINs progress to PDAC, laying a basis for future early detection of this and different sorts of pancreatic most cancers.
The three-way evaluation revealed that some key options of pancreatic most cancers had been current in PanINs.
“When we appeared on the development of regular to excessive grade PanIN lesions, we discovered that cell proliferation regularly will increase whereas inflammatory signaling decreases, which will be necessary to know the intrinsic low immunogenicity of those tumors,” says Kagohara. This means that the cells in and round PanINs are already making a extra immunosuppressive setting earlier than the invasive PDAC is totally developed, she says.
The findings additionally revealed spatial variations within the cells. “We discovered that cancer-associated fibroblasts, which play a serious position within the biology of PDAC and their response to remedies, had been already current on the premalignant stage, per earlier analysis in animal fashions of pancreatic most cancers however not beforehand noticed in people,” says Kagohara.
“PanINs are fairly small—lower than 1 millimeter in dimension—so we had been very stunned that even utilizing only a few cells, we had been capable of detect sturdy signatures from these lesions,” she provides.
“That’s crucial in most cancers as a result of, for instance, if we wish to perceive immune responses to remedies, we have to know which immune cells are nearer to or farther from the tumor. But we additionally want to know if there are different cell sorts which can be blocking these communications by forming a bodily barrier or interacting with the neoplastic cells.”
Beyond the primary findings, the research highlights the facility of computational instruments to additional understanding of human illness.
Spatial transcriptomics is a really highly effective know-how for producing information, says Kagohara, however the precise computational instruments are wanted to investigate it. “With the precise instruments, you may improve the outcomes which you could extract from these novel applied sciences,” she says.
To encourage extra collaboration and analyses, the research’s information is offered on GEO (GSE254829), and the code is offered on GitHub.
“As we generate an increasing number of information on PanINs, the concept is to determine molecular signatures for the event of early detection assessments,” says Kagohara. “This is among the first research to generate a useful resource open to different researchers within the seek for these early markers in PanINs so we are able to study extra about how the spatial distribution of molecular and mobile signatures have an effect on the development of pancreatic most cancers and the response to remedies.”
Other research co-authors embrace Alexander Bell, Jacob Mitchell, Ashley Kiemen, Melissa Lyman, Kohei Fujikura, Jae Lee, Eron Coyne, Sarah Shin, Sushma Nagaraj, Atul Deshpande, Pei-Hsun Wu, Dimitrios Sidiropoulos, Rossin Erbe, James Chell, Lauren Ciotti, Jacquelyn Zimmerman, Denis Wirtz, Won Jin Ho, Neeha Zaidi, Elizabeth Thompson, Elizabeth Jaffee and Laura Wood of Johns Hopkins. Researchers from 10x Genomics of Pleasanton, California, contributed to the work.
More data:
Alexander T.F. Bell et al, PanIN and CAF transitions in pancreatic carcinogenesis revealed with spatial information integration, Cell Systems (2024). DOI: 10.1016/j.cels.2024.07.001
Johns Hopkins University
Citation:
Identifying key markers in pancreatic most cancers development utilizing a brand new evaluation pipeline (2024, August 29)
retrieved 29 August 2024
from
This doc is topic to copyright. Apart from any truthful dealing for the aim of personal research or analysis, no
half could also be reproduced with out the written permission. The content material is offered for data functions solely.